COSMIC Release v88
19 Mar 2019
The COSMIC v88 release is now live! In this release we have 4 newly expert-curated genes: PTPRT, MAP3K1, LEF1 and LATS2; and a significant update to TSC1 and TSC2. We've incorporated 205 whole genome screened samples from 10 systematic screen papers. We've also added 17 new genes that have Hallmarks of Cancer.
Importantly, we have also updated the HGVS syntax on some of the manually curated mutations in COSMIC. This is one of the steps we will be taking towards the extensive reannotation of the entire COSMIC database. Further significant changes will be coming in future releases. For details of the HGVS changes you can expect, please see the help documentation.
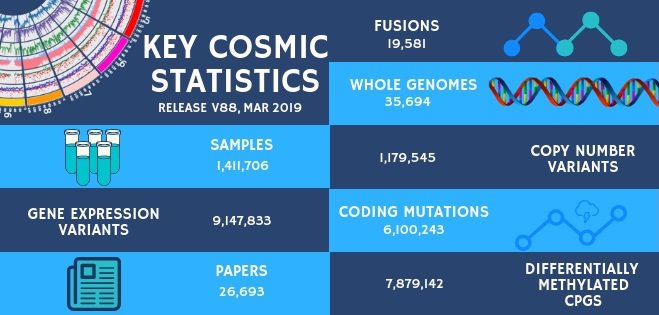

New genes: PTPRT (protein tyrosine phosphatase receptor-type, T) encodes a protein tyrosine phosphatase (PTP) which is a member of the R2B subfamily of the PTP family of proteins. Located on chromosome 20 (20q13.11), PTPRT is the most frequently mutated PTP gene in human cancer and genetic changes have been observed in a variety of cancer types including colon, endometrial, bladder, oesophageal, head and neck, lung, skin and stomach. These mutations are mainly missense single nucleotide variants, distributed throughout the gene, although nonsense, insertion and deletion mutations that result in premature truncation of the protein have also been found. This distribution of mutation types and sites is suggestive of a tumour suppressor gene. An idea that is further supported by studies in knock-out mice and the fact that loss of PTPRT function also occurs in human cancers through methylation of promotor DNA. PTPRT is thought to act as a tumour suppressor by dephosphorylating activated STAT3 and paxillin, two substrates which have an oncogenic role in tumour development, when phosphorylated. PTPRT also has a role in cell-cell adhesion which is affected negatively by loss of function mutations. In addition, deleterious mutations in PTPRT (predominantly those occurring in the phosphatase domain) have been implicated in the drug resistance mechanism of metastatic colorectal carcinomas treated with bevacizumab together with chemotherapy.
MAP3K1 (mitogen-activated protein kinase kinase kinase 1) encodes a protein which is a serine/threonine kinase and part of some signal transduction cascades, including the ERK and JNK kinase pathways as well as the NF-kappa-B pathway. Recurrent loss of function MAP3K1 mutations, including frameshifts, nonsense and splicing mutations, suggesting a tumour suppressor role, have been found in breast cancer. The mutations are most frequent in the luminal A subtype and are associated with indolent disease and a favourable prognosis. Genome studies have also identified low to moderate frequency MAP3K1 deletions and mutations in other tumour types such endometrial, colorectal and lung.
LEF1 (Lymphoid enhancer factor 1) located at 4q25, encodes a member of the lymphoid enhancer factor/T-cell factor (LEF/TCF) family of DNA binding transcription factors and shares homology with the high mobility group protein-1. LEF1 interacts with nuclear β-catenin in the Wnt signalling pathway and can act as either a tumour suppressor gene or an oncogene. Two major isoforms are found, including a short form which lacks the β-catenin binding domain at the N terminus and shows dominant negative activity, suppressing the Wnt pathway by preventing β-catenin recruitment. LEF1 plays an important role in early lymphoid development and increased expression is associated with poor prognosis in several leukaemias. Somatic mutations and deletions/microdeletions have been found in a variety of childhood and adult T and B cell acute lymphoblastic leukaemias (ALL). These mutations include truncating frameshift and nonsense mutations as well as gain-of-function missense mutations in the β-catenin binding domain, and these missense mutations have been shown to be associated with proliferation of ALL cells. Expression of N terminus-truncated LEF1 in mice results in tumours with sebaceous differentiation, and in humans nonsense and missense mutations located in the β-catenin binding domain which result in inactivation of Wnt signalling have been found in sebaceous carcinomas and adenomas as well as benign sebaceomas.
LATS2 (large tumour suppressor kinase 2), found at the genomic locus 13q12.11, encodes for a serine/threonine kinase which acts in the Hippo signalling pathway via YAP/TAZ proteins by restricting proliferation and promoting apoptosis. Its role in carcinogenesis is mediated in part via somatic mutations in many tumour types. Mutations such as nonsense, frameshift and missense mutations are likely to affect LATS2 functionality, possibly through negatively affecting interactions with positive regulators such as NF2 or MOB1, by damaging the catalytic domain of LATS2, interfering with activating phosphorylations through upstream kinases, or other mechanisms. Data from malignant mesothelioma indicate that Hippo pathway dysregulation is frequent in MM cells with inactivation of LATS2 or an upstream regulator, such as NF2, of this pathway. In a series of malignant pleural mesothelioma patients the LATS2 gene was altered in 11% of MPM by point mutations and large exon deletions. Genetic data coupled with transcriptomic data allowed the identification of a new MPM molecular subgroup, C2LN, characterized by a co-occurring mutation in the LATS2 and NF2 genes in the same MPM. MPM patients of this subgroup presented a poor prognosis.
Update to TSC1 and TSC2, Tuberous sclerosis complex (TSC) is an autosomal dominant disorder characterized by skin manifestations and formation of multiple tumours in different organs. In addition, TSC disease causes disabling neurological disorders, including autism, intellectual impairment and seizures. TSC1 and TSC2 encode proteins hamartin and tuberin, respectively, which interact and form a heterodimer that inhibits the activation of mammalian target of rapamycin (mTOR) complex 1 (mTORC1), a master regulator of nutrient and growth factor induced signalling. Tumorigenesis involves inactivation of the second allele, typically via a second-hit somatic mutation in the wild-type allele of the TSC gene. Current COSMIC release reports somatic mutations in renal cell carcinoma (RCC), subependymal giant cell astrocytomas and other gliomas, bladder cancer, hepatocellular carcinoma, melanoma, angiomyolipoma and lymphangioleiomyomatosis amongst many other tumour types. Full details of this curation can be seen here.
In addition to the curated genes, 17 new genes have been added to the Hallmarks of Cancer. These are BCL9L, MAP3K13, TRAF7, CTNNB1, NT5C2, UBR5, FBXO11, PBRM1, ZRSR2, GRIN2A, PLCG1, H3F3B, POT1, KMT2C, SIX1, KMT2D, and STAG2. There are 10 new systematic screens in v88, for further technical details please see the release notes.
